Read our Privacy Notice if you are concerned with your privacy and how we handle personal information. If you plan to use these services during a course please contact us. You can choose the genomic sequence strand that may be ‘Plus’, ‘Minus’ or ‘Both’. The General Options tab allows to set various options. Here you can select multiple genomic ranges or transcripts to be aligned to one protein. If you have any feedback or encountered any issues please let us know via EMBL-EBI Support. ProSplign generates pairwise alignment between a protein and a genomic sequence. Please read the provided Help & Documentation and FAQs before seeking help from our support staff. The tools described on this page are provided using Search and sequence analysis tools services from EMBL-EBI in 2022 GeneWise compares a protein sequence to a genomic DNA sequence, allowing for introns and frameshifting errors. Genomic alignment tools concentrate on DNA (or to DNA) alignments while accounting for characteristics present in genomic data. SSEARCH2SEQ finds an optimal local alignment using the Smith-Waterman algorithm. LALIGN finds internal duplications by calculating non-intersecting local alignments of protein or DNA sequences. 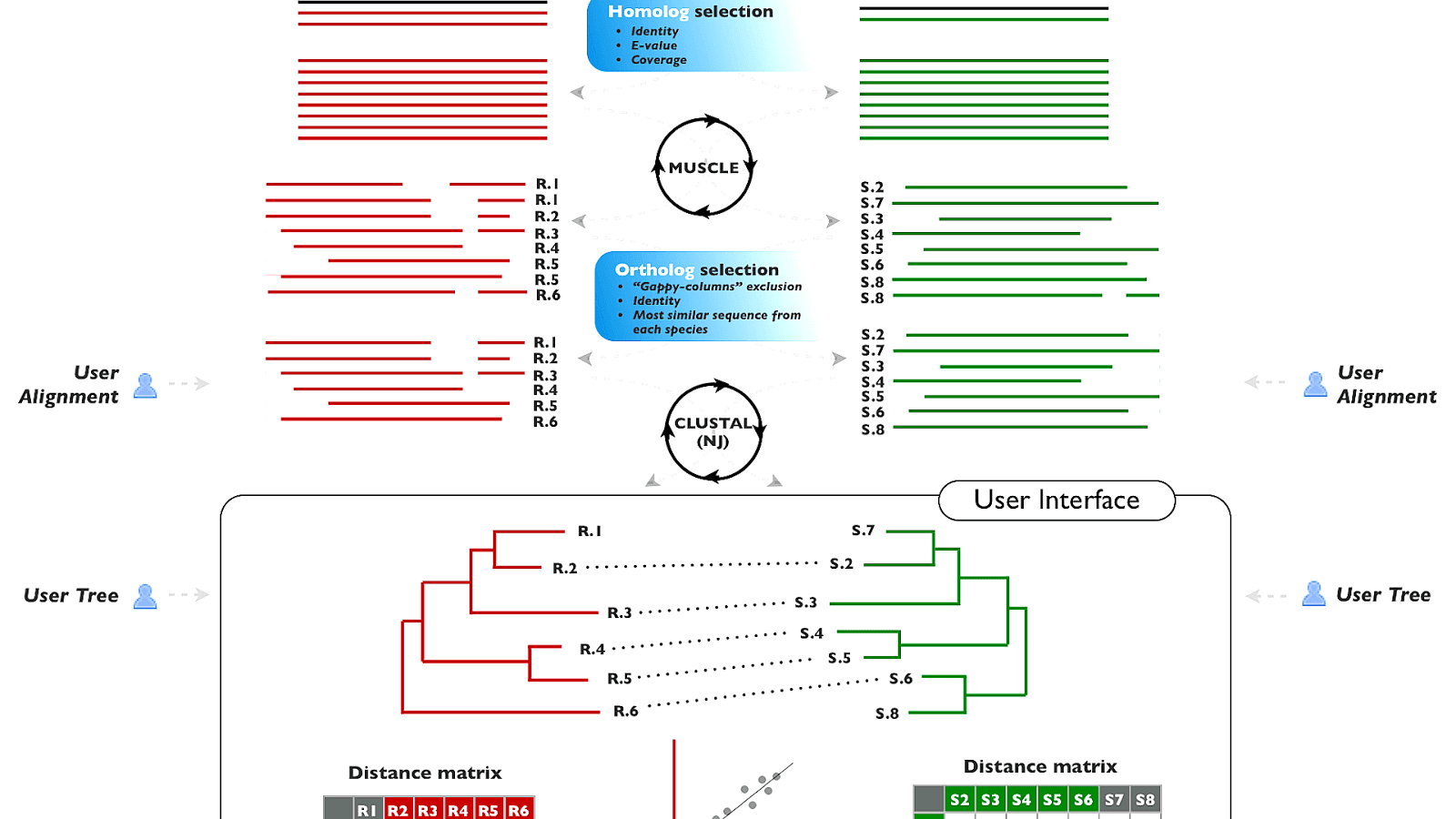
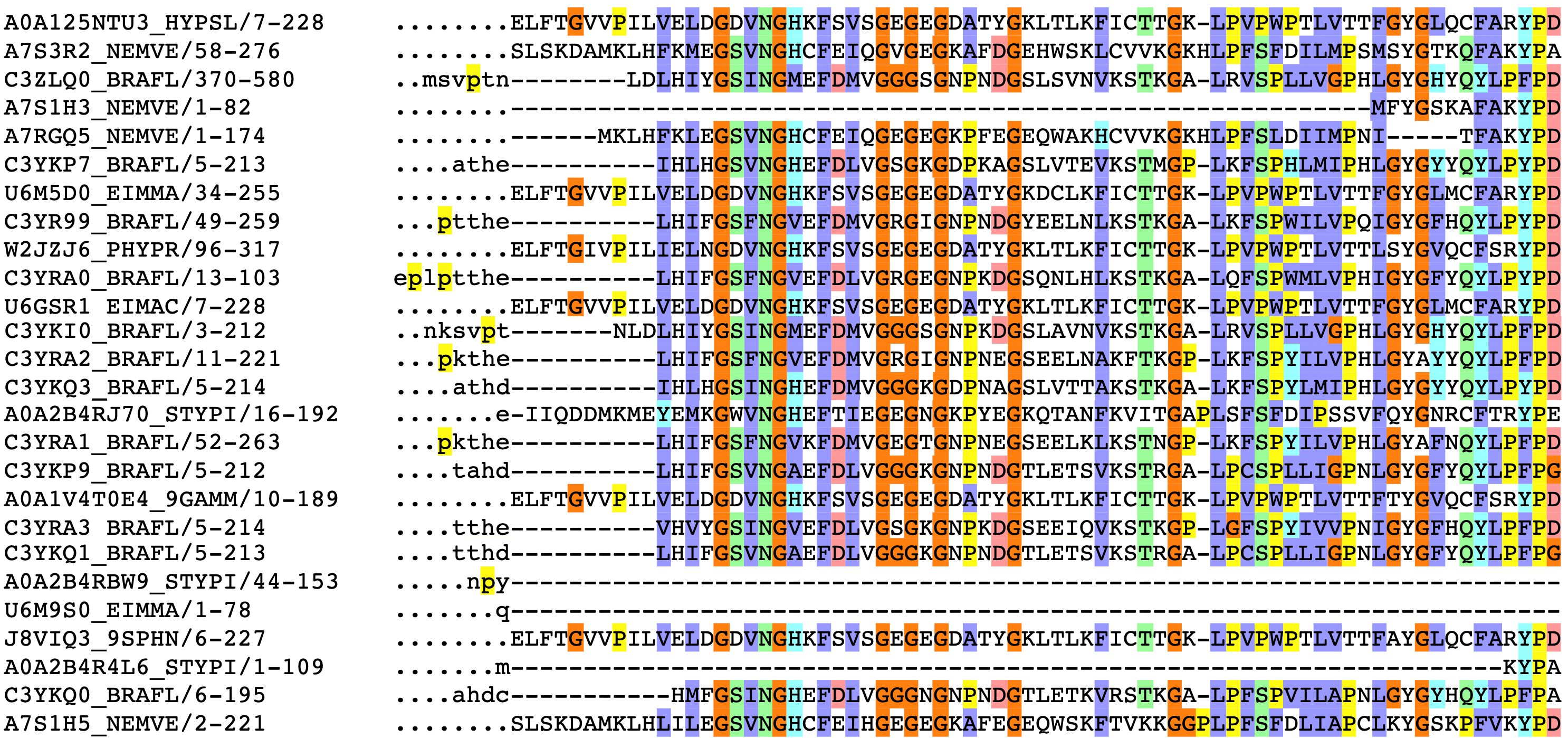
They are can align protein and nucleotide sequences.ĮMBOSS Water uses the Smith-Waterman algorithm (modified for speed enhancements) to calculate the local alignment of two sequences.ĮMBOSS Matcher identifies local similarities between two sequences using a rigorous algorithm based on the LALIGN application. Local alignment tools find one, or more, alignments describing the most similar region(s) within the sequences to be aligned. GGSEARCH2SEQ finds an optimal global alignment using the Needleman-Wunsch algorithm. Biopython has a wide range of functionalities for. Identification of similar provides a lot of information about what traits are conserved among species, how much close are different species genetically, how species evolve, etc. Global alignment tools create an end-to-end alignment of the sequences to be aligned.ĮMBOSS Needle creates an optimal global alignment of two sequences using the Needleman-Wunsch algorithm.ĮMBOSS Stretcher uses a modification of the Needleman-Wunsch algorithm that allows larger sequences to be globally aligned. Sequence alignment is a process in which two or more DNA, RNA or Protein sequences are arranged in order specifically to identify the region of similarity among them. From the output of MSA applications, homology can be inferred and the evolutionary relationship between the sequences studied. Pairwise Sequence Alignment is used to identify regions of similarity that may indicate functional, structural and/or evolutionary relationships between two biological sequences (protein or nucleic acid).īy contrast, Multiple Sequence Alignment (MSA) is the alignment of three or more biological sequences of similar length. If two-protein families were also taken into account, extHomFam would be increased to 1013 sets, at the cost of positively biasing the TC score (with two reference sequences it becomes equal to SP.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
